S Schirmer, F Langbein, C Jenkins, M Chandler. Design of novel MRI pulse sequences for GABA quantification using optimal control. In: 4th Int Symp on MRS of GABA, 2017.
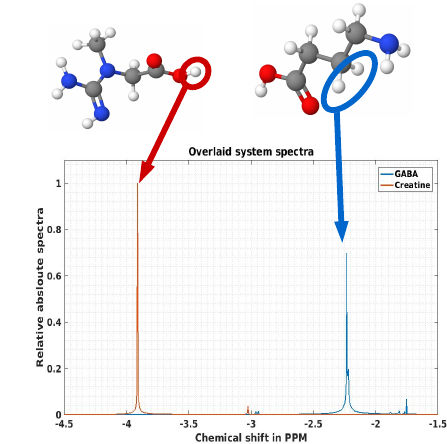
Discrimination and quantification of chemicals such as GABA, Glutamine,Glutamate, based on in vivo spectra obtained with clinical MRI systems, is complicated by overlapping spectra of other molecules and a range of uncertainties during acquisition. Several approaches to address this problem have been developed, including spectral editing (MEGAPRESS) and 2Dspectroscopy. An alternative approach that promises greater flexibility and robustness is the automated design of selective pulse sequences for molecular biomarkers such as GABA using quantum control techniques. Recent successful applications of quantum control to optimize NMR pulse sequences strongly support this approach. Computational results demonstrate that optimal pulse sequences, in principle, generate clear spectra with distinct, easily identifiable features for metabolites that ordinarily have overlapping spectral features. While there are trade-offs between selectivity and robustness,simulations suggest that they can be made robust with regard to B0inhomogeneity, flip angle errors and noise. However, implementation of such optimal control pulses faces several challenges, including the complexity of optimal pulses compared to standard pulse sequences and a lack of sufficiently accurate NMR models for molecules in vivo. Hence, optimization and testing of chemically selective pulse sequences requires tissue-mimicking phantoms with known and accurately controllable chemical compositions to construct accurate models for auto-optimization and validate pulse sequence performance. We discuss preliminary robust optimal control designs based on NMR models of metabolites and tissue-mimicking phantom results.
![]() This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.